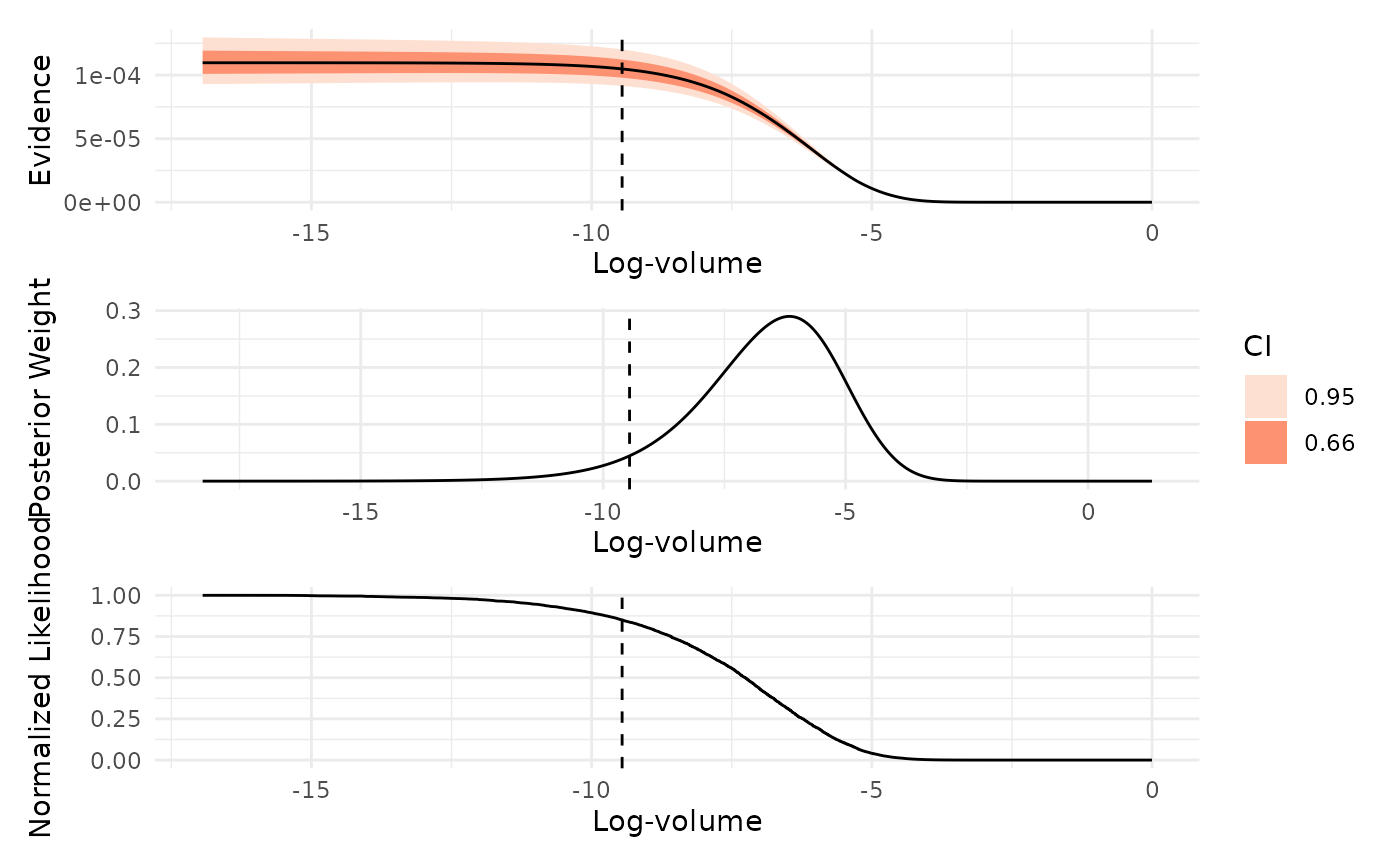
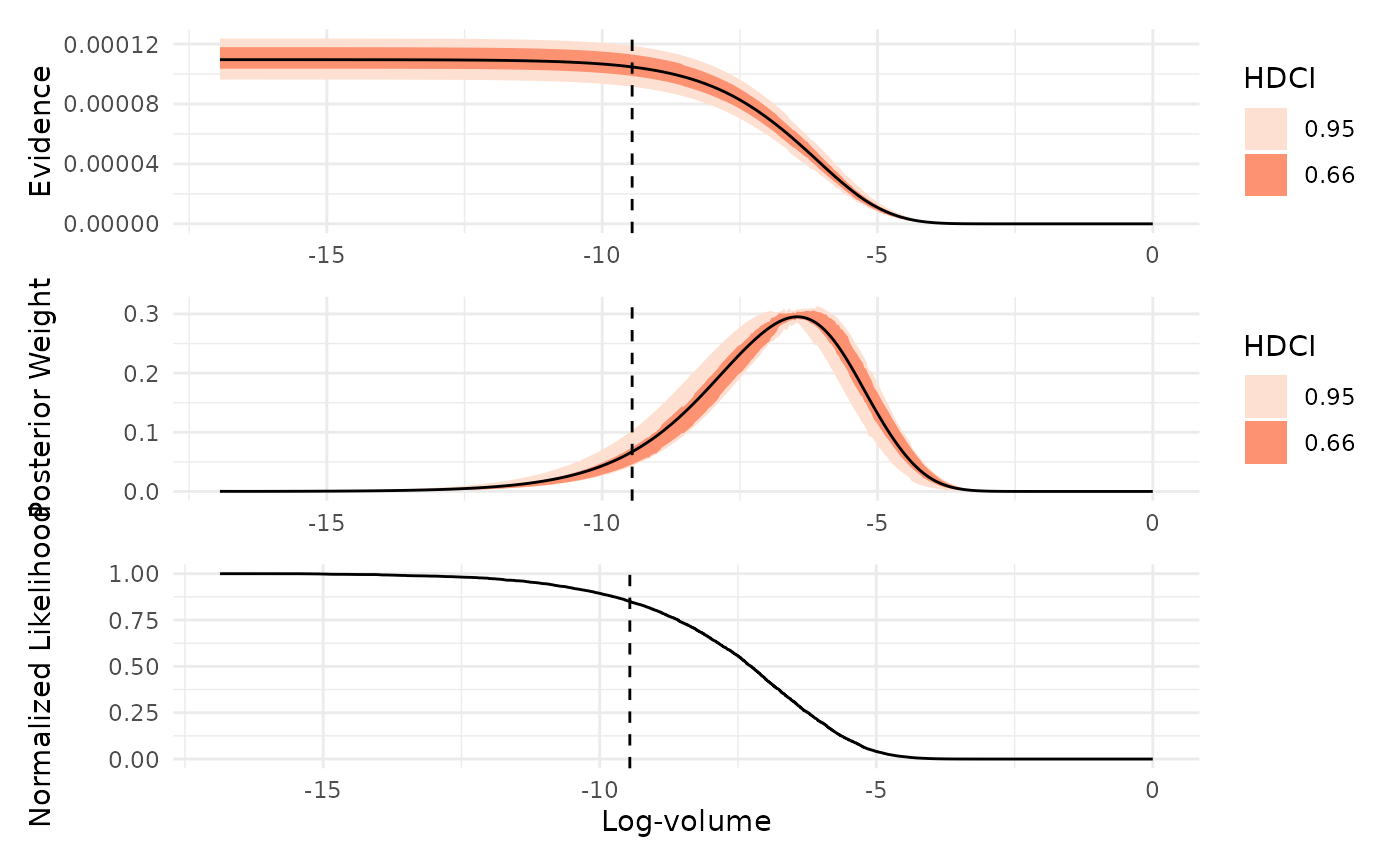
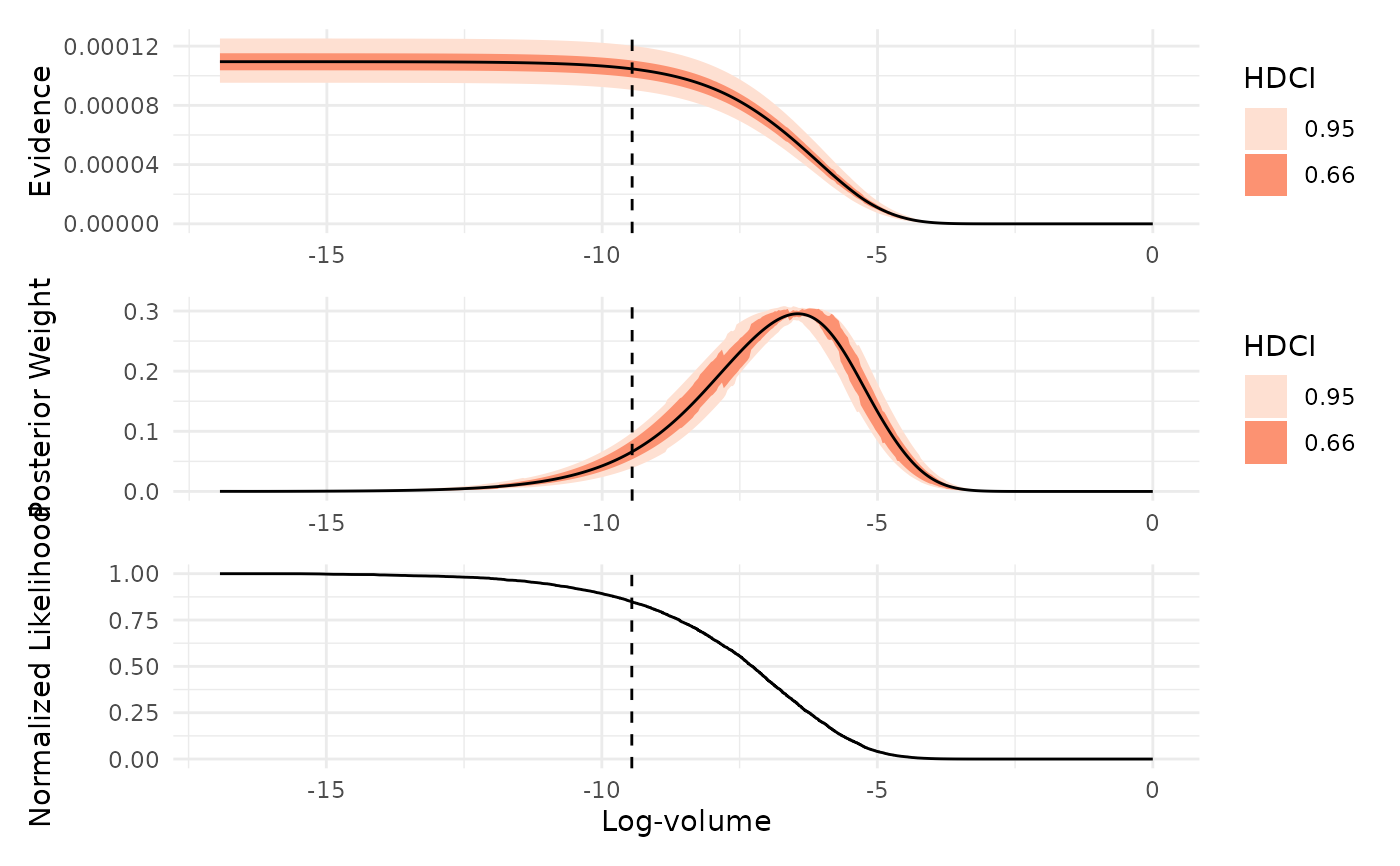
Visualizes key diagnostics from a nested sampling run, including the normalized likelihood, posterior weights, and evidence as functions of log-prior volume.
Arguments
- x
[ernest_run] or [ernest_estimate]
An object containing results from nested sampling.- which
[character()]
Choose which plots to display. Must be one or more of"evidence","weight", and"likelihood".- ...
These dots are for future extensions and must be empty.
- ndraws
[integer(1)]
The number of log-volume sequences to simulate. If equal to zero, no simulations will be made, and a one draw vector of log-volumes are produced from the estimates contained inx.
Value
[invisible(x)]. A ggplot2::ggplot() object is printed as a
side effect.
Details
Interpreting these plots can help diagnose issues such as poor or
insufficient prior sampling and model misspecification. Use which to select
the plots to display:
which = "evidence": Plots the estimated marginal likelihood (evidence) as a function of log-prior volume, with uncertainty intervals. Peaks in this plot indicate regions of prior volume that contribute most to the evidence estimate.which = "weight": Shows the distribution of posterior mass across log-prior volume. This plot highlights which regions of the prior volume contain the most posterior probability, helping to identify where the sampler concentrated its effort.which = "likelihood": Displays the normalized likelihood as a function of log-prior volume. Smoothness in this plot indicates effective likelihood-restricted prior sampling, while irregularities may suggest sampling difficulties or, in some cases, misspecified likelihood functions.
If x is an ernest_run, the plots are based on the actual run data. Error
ribbons are drawn around the evidence plot from analytic estimates of
uncertainty (see summary.ernest_run).
If x is an ernest_estimate (or if ndraws is specified), the plots
are based on simulated values from the log-volume. The highest density
continuous intervals (HDCIs) are computed using ggdist::median_hdci() for
both the evidence and weight plots.
Note
Plotting multiple diagnostics with which requires the patchwork package.
Plotting ernest_estimate objects requires the ggdist package.
See also
calculate()for generatingernest_estimateobjects.visualize()for plotting the posterior distributions generated by a run.